|
I think you’ll see that violin plots are a little more involved to create, and they are slower to render, but many find them to be more informative and more aesthetically pleasing. We’ll neeed to load a few packages to add functions to help us. Violin plots are similar to boxplots except that they attempt to present the distribution of the data that would otherwise be represented by a box. Here we’ll demonstrate the use of barplots and violin plots to visualize this data. We see that it is a large matrix containing lots of numbers, so viewing it directly is of little use. # browseVignettes('vcfR') # Documentation We can take a quick peak at this to remind us what it looks like. We can extract the depth data (DP) as explained elsewhere. Here we visualize the depth information from a VCF file to provide a perspective on sequence quality. Make sure to check your variant caller documentation if you do not see this information in your file, there is a chance it was an option that was not selected. Many variant callers report this information, but not all. 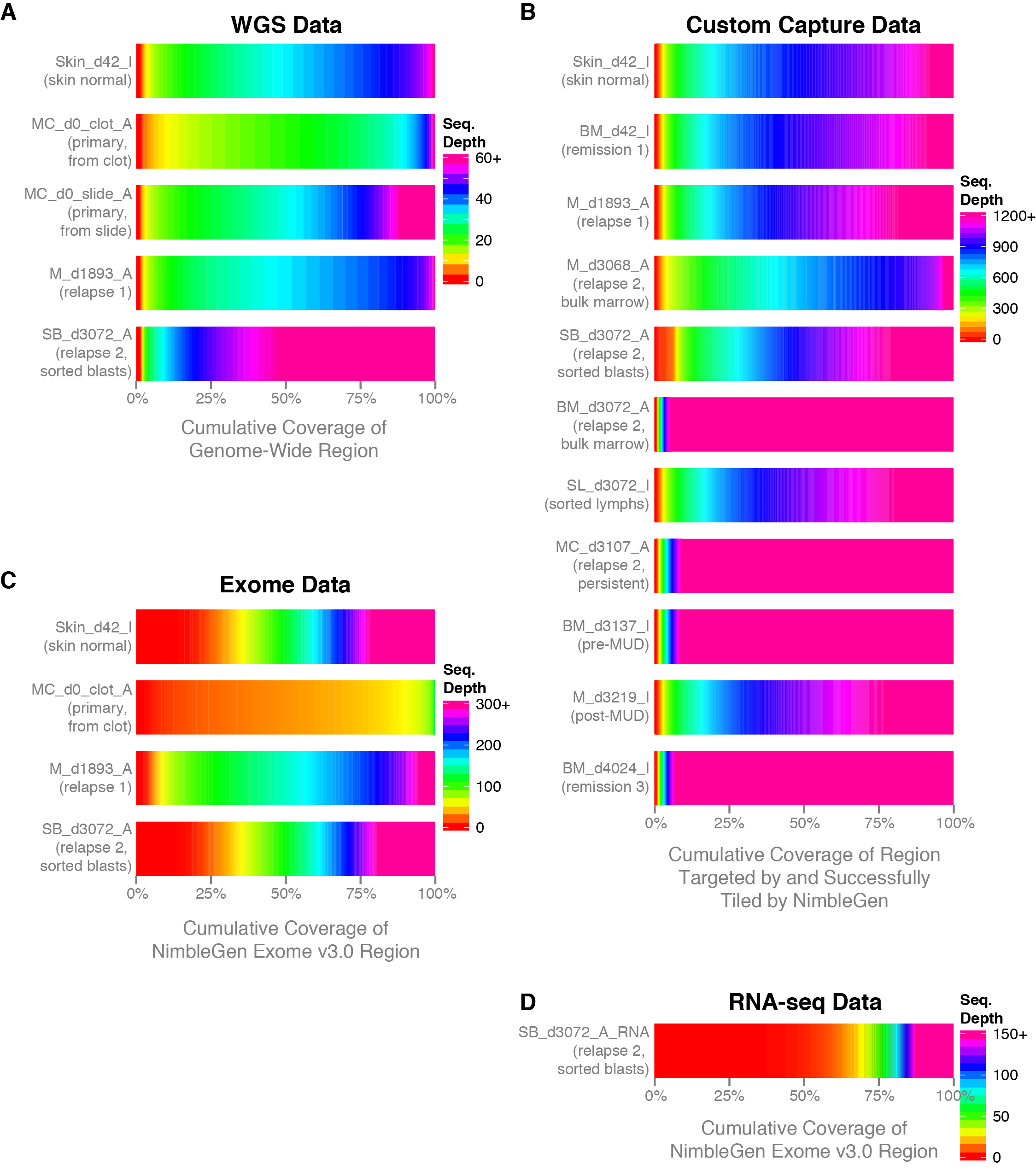
But this can be used as a convenient subset of an entire genome. In VCF data only information on the variable positions are reported. At this point one of the first questions asked is how well did the sequencing go? One measure of sequencing quality is sequence depth or how many times each position was sequenced. Once a sequencing run is complete the data are typically mapped to a reference genome, variants are called and these variant are stored in a VCF file.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
